AbstractIn this review of the most recent applications of deep learning to ultrasound imaging, the architectures of deep learning networks are briefly explained for the medical imaging applications of classification, detection, segmentation, and generation. Ultrasonography applications for image processing and diagnosis are then reviewed and summarized, along with some representative imaging studies of the breast, thyroid, heart, kidney, liver, and fetal head. Efforts towards workflow enhancement are also reviewed, with an emphasis on view recognition, scanning guide, image quality assessment, and quantification and measurement. Finally some future prospects are presented regarding image quality enhancement, diagnostic support, and improvements in workflow efficiency, along with remarks on hurdles, benefits, and necessary collaborations.
Deep learning, as a representative technology in the field of artificial intelligence (AI), has already brought about many meaningful changes in ultrasonography [1,2]. The tremendous potential of this technology, both clinically and commercially, is widely recognized in academia and industry. This new trend is a leap forwards from the traditional ultrasound technology inspired by information technology (IT) and consumer electronics technology. New AI-based applications range from enhancing ultrasound images [3-8] to smart and efficient improvements of the workflow of healthcare professionals [9-18]. Deep learning applied to imaging chains has shown major improvements in the efficiency and effects of processing, such as signal acquisition, adaptive beamforming, clutter suppression, and compressive encoding for color Doppler [19]. It has also been shown that deep learning implementation in standard clinical fields, such as breast and thyroid ultrasound imaging, could increase diagnostic accuracy, reduce medical costs, and provoke some insightful discussions. In addition, using data augmentation could improve the generalizability of deep learning models, and introducing a transparent deep learning model to explain how and why AI systems make predictions could build trust in AI systems [20].
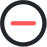
Enthusiasm for this technology can be easily demonstrated by the number of publications. For example, as seen in Fig. 1, the number of deep learning papers in PubMed soared since 2017 to more than 4,000 in 2019, and the number of deep learning applications in ultrasonography also followed the same trend. The wide participation from academia, clinical institutions, and industry is a clear indicator of eagerness and expectations for this technology.
This review paper will briefly introduce deep learning technology and major components thereof in ultrasound applications, and summarize the practical applications of deep learning in ultrasonography (especially in the domains of imaging, diagnosis, and workflow), focusing on the most recent research. A short discussion of future opportunities will also be presented.
Progress in algorithms, improved computing power, and the availability of large-scale datasets are the three major components responsible for the recent success of deep learning technology [21]. Many competitively developed algorithms are being updated and made accessible in deep learning frameworks such as Caffe [22], TensorFlow [23], Keras [24], PyTorch [25], and MXNet [26].
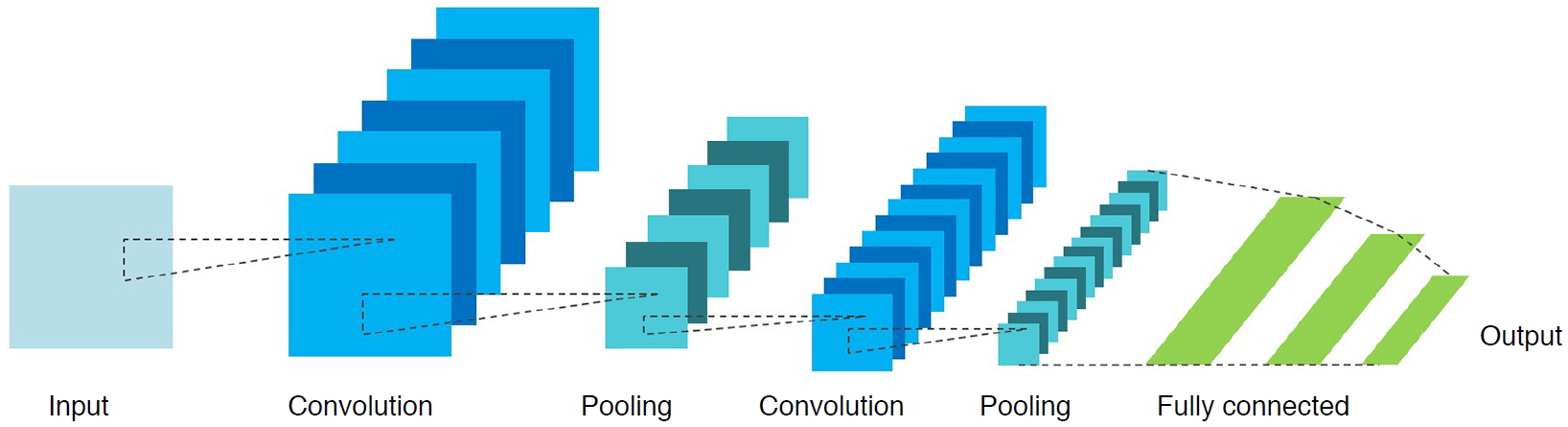
Convolutional neural networks (CNNs) have played the most important role in the history of adoption of deep learning for video and image processing applications. As illustrated in Fig. 2, a CNN with a feature map extracting the convolutional layer and size-reducing pooling layer automatically extracts the optimized output through training process, thereby expediting applications in imaging. CNNs can be utilized for various purposes depending on their structure and training data. Concerning output features in scanned images can be recognized and automatically assigned to a meaningful category (classification). A specific feature or object can be located (detection), and the edge of an object can be precisely delineated (segmentation). Furthermore, new images, not visibly distinct from real ones, can be fabricated (generation).
CNNs have been expanded from natural image classification networks including AlexNet [27], VGGNet [28], GoogLeNet [29], ResNet [30], and DenseNet [31]. Key components of these developments include deeper networks with more convolutions per layer, and adoption of new layers, such as skip connections, to deliver information to deeper layers. The basic architecture of CNNs for classification is shown in Fig. 3.
Object detection is a method for recognizing the type and location of an object in an image. Object detection methods using CNNs are broadly divided into two types. In two-stage detection, a region of interest (ROI) is detected by a region proposal and classification is then performed of the ROI, as shown in Fig. 4A. The region proposal finds possible locations for objects. Detection can be also implemented in one stage, such that ROI detection and classification are performed simultaneously, as shown in Fig. 4B.
Segmentation divides an image according to a rule reflecting the question of interest with resolution meaningful for applications. Each pixel of the image is first classified and then de-convoluted to compensate for the pooling. Fig. 5 presents a typical architecture of U-net [36] based segmentation. Representative networks for segmentation include fully convolutional networks [37] and DeconvolutionNet [38], and various U-Net-based networks [38-41] have been proposed.
The structure shown in Fig. 6 can be used to generate new unseen images based on the given images that originally existed. A generative adversarial network (GAN) can train and generate such images in a way that the generator and the discriminator compete within the network; the generator generates various new images from the random variables, and the discriminator distinguishes whether the images are real or generated. A sufficiently trained GAN can produce images that are not distinguishable from the real ones. This feature provides several applications, including augmentation of training data and image quality and resolution improvements [42,43]. Fig. 7 shows some examples of classification, detection, segmentation, and generation as applied to ultrasound images.
Ultrasound signals and images have unique characteristics and issues not as strongly present in other imaging modalities, such as attenuation, penetration, uniformity, shadowing, real time, and operator-dependence. These specific aspects must be taken into account when applying deep learning to ultrasonography. That means that a careful understanding of the system, its usage, and the practice environment should precede the design and implementation of a deep learning-based system. Most ultrasound practice is performed in real time, which in turn requires real-time output for deep learning-based functions. Additionally, the vast array of transducers, settings, and scan modes require corresponding diversity and integrations.
It is necessary to establish a standardized training data set because ultrasound images have strong operator-dependence and different image characteristics for each device. There are standardized imaging guidelines for each clinical scan, but when training and using a deep learning model, it is necessary to precisely define in advance the section of the ultrasound images before acquiring them. Additionally, pre-processing and normalization may be required to remove non-image information, such as annotations, and to reduce image deviation due to various scan conditions, respectively. The size of the data available for training is also an important consideration. The issue of small data sets can be alleviated by using transfer learning [44] where, as shown in Fig. 8, a pretrained model based on a large dataset can be effectively used in a smaller dataset when the two datasets share certain low-level features such as edges, shapes, corners, and intensity. For example, transfer learning enables us to utilize the knowledge of a pretrained breast lesion detection model trained with a large number of breast images to train a model on thyroid lesion detection with a smaller number of thyroid images.
The adoption of deep learning in ultrasound imaging can be explained from a simplified perspective, as shown in Fig. 9. For the sake of convenience of discussion, the task can be divided into the domains of medical practitioners, ultrasound imaging systems, and deep learning engines. Scanned images are processed by an ultrasound imaging system to produce output images, of which measurement and/or quantification are then performed. Assistance can be provided in scanning by automatic recognition of which organ is being scanned, guidelines on how to scan, and assessments of scanned image quality. Traditional signal processing can be further enhanced in areas including beamforming, higher resolution, and image enhancement. The laborious and repetitive job of measurement/quantification can be replaced by computer-aided detection (CADe) and computer-aided quantification (CADq). Finally, physicians can consult second opinions from computer-aided diagnosis (CADx) and/or computer-aided triage (CADt) systems.
Image processing has been enriched by deep learning, opening up a vast array of opportunities and improvements. Conventional signal processing is being combined with deep learning to produce better images [4,45], methods previously deemed to be practically unfeasible are being implemented [4,46], computation time is being greatly reduced [4,46], and new images can be created from the scanned images [47].
Yoon et al. [4] presented a framework for generating B-mode images with reduced speckle noise by using deep learning. The proposed method greatly exceeded the ability of the traditional delay-and-sum (DAS) beamformer, while maintaining the resolution. Luijten et al. [46] showed that a content-adaptive beamformer, such as an eigenspace-based minimum variance (MV) beamformer, which had not previously been utilized for real time applications, was implementable though training. The processing time of the MV beamformer, typically 160 seconds per image, was reduced to 0.4 seconds per image with similar image acquisitions. It is expected that new beamformers performing better than conventional DAS will be implemented in the near future [4,46,48-50].
Liu et al. [47] showed the future potential of ultrasound localization microscopy (ULM) by implementing U-Net based ULM. The system could detect micro-bubbles (17 μm), taking about 23 seconds per image; this was still a long time, but several times faster than the previous version, indicating the possible future applicability of this system. Huang et al. [51] proposed MimicKNet, which imitates the post-processing technique for ultrasound based on GAN. Based on DAS and taking logarithms of the data, it was trained on 1,500 cine loops with 39,200 frames of fetal, phantom, and liver targets, and was applied to untrained frames of the heart to show the capability of real-time processing of 142 frames per second using a P100 GPU.
Jafari et al. [52] provided a deep learning solution that modified the low-quality images from a point-of-care ultrasound (POCUS) device to a level of quality comparable to the that obtained using premium equipment. By employing constrained CycleGAN, the experiment could also improve the accuracy of automatic segmentation using POCUS data. Wildeboer et al. [53] presented methods of generating synthesized shear-wave elastography (SWE) images based on original B-mode images. Using both B-mode and SWE images collected from 50 prostate cancer patients, it was shown that synthesized SWE images with an average absolute error of pixel units within 4.5±0.96 kPa could be generated.
The deep learning application with the most fundamental effect on ultrasound imaging is ultrasonic beamforming. The DAS beamformer, most widely used beamforming method in ultrasound systems, has become an industry standard because it can be applied in real-time with a small amount of computation. However, the DAS algorithm utilizes predefined apodization weights, leading to low-resolution images with strong artifacts and poor contrast due to high sidelobes. A variety of adaptive beamforming methods have been still proposed to address the shortcomings of DAS beamforming, but it remains difficult to deem any of them clinically meaningful for general purposes due to their computational complexity. Recently, however, as the limitations of these computations are being overcome by using deep learning, the possibility that adaptive beamforming could be performed in real time is emerging [45,46].
CADx assists doctors to improve diagnostic accuracy and consistency by suggesting second opinions. Deep learning-based CADx is expanding rapidly, covering more organs and diseases in many imaging modalities. From conventional machine learning methods, where manually selected features were utilized, especially in ultrasonography, deep learning-based studies are moving towards multi-parameter and multi-modality fusion of various information including non-ultrasound imaging, clinical information, and genotype information. In ultrasonography itself, accuracy can be improved by using various types of information, such as Doppler and elastography, other than B-mode [54,55].
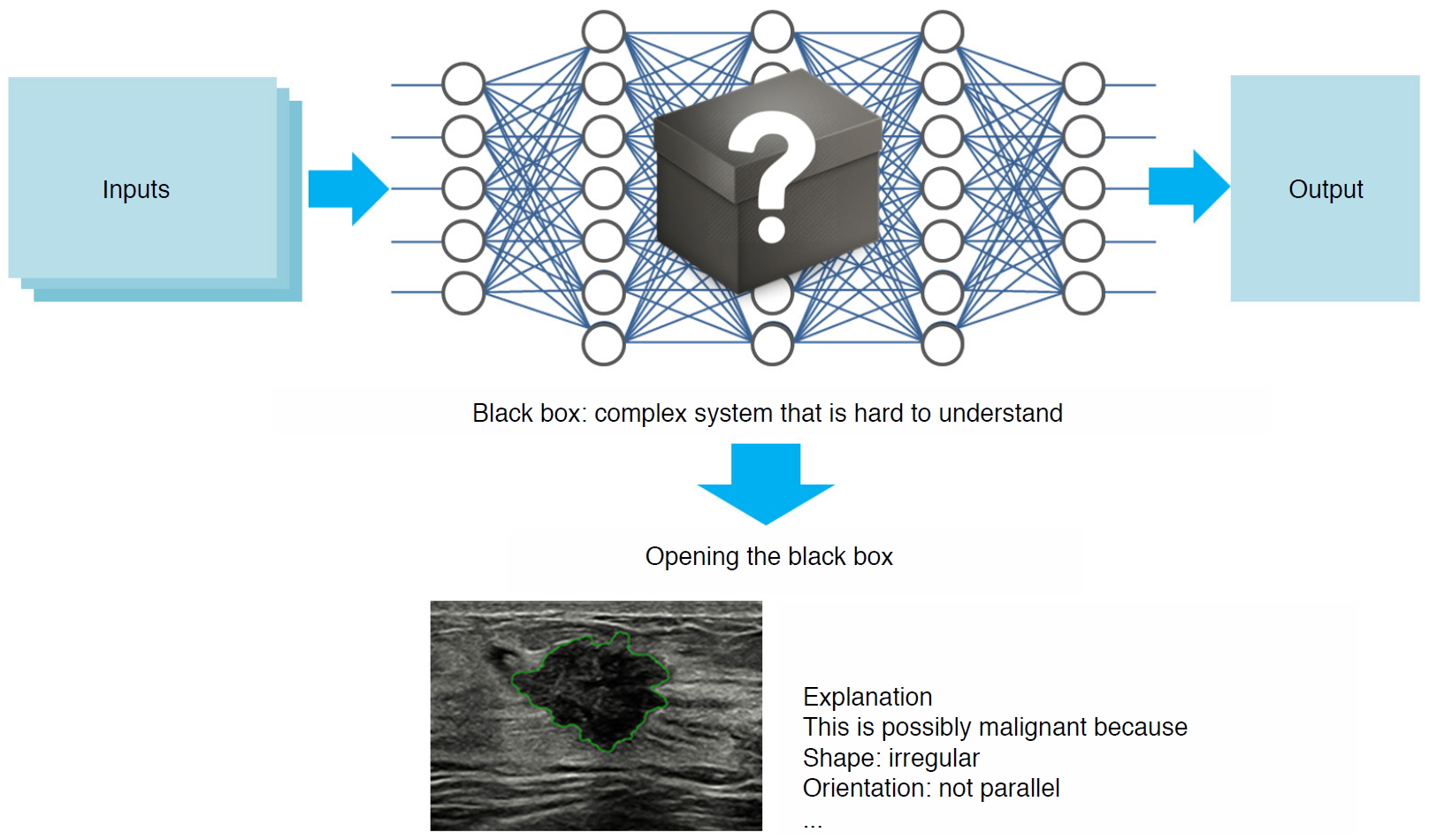
Another important topic in diagnosis is eXplainable AI (XAI). As illustrated in Fig. 10, deep learning technology has been regarded as just a black box that cannot interpret the process of deriving the output. However, Ghorbani et al. [56] have shown that a CNN applied to echocardiography can identify local cardiac structures and provide interpretations by visually highlighting hypothesis-generating regions of interest. As such, studies published in the field of XAI have sought to provide explanations of why deep learning models produce certain outputs. Another issue to be addressed when discussing XAI is that there is no analytical explanation of what mechanisms it operates through, except for the explanation that a deep learning model works because it has found optimal parameters through training with big data. Ye et al. [57] demonstrated that the success of deep learning stems from the power of a novel signal representation using a nonlocal basis combined with a data-driven local basis, which is indeed a natural extension of classical signal processing theory.
Breast cancer is the most common cancer in women. As noted in Table 1, diverse studies with deep learning applications are being conducted. Early studies only utilized B-mode images, but recent studies have concentrated on combining the usage of ultrasound multi-parametric images or clinical information. Zheng et al. [58] introduced a new method to determine metastasis to axillary lymph nodes in early-stage breast cancer patients. Features obtained from deep learning-based radiomics were combined with clinical parameters such as patient age, size of the lesion, Breast Imaging Reporting and Data System (BI-RADS) category, tumor type, estrogen receptor status, progesterone receptor status, human epidermal growth factor receptor 2 (HER2), Ki-67 proliferation index, and others. Sun et al. [59] included additional molecular subtype information such as HER2-positivity and triple-negative status. Luminal and Liao et al. [54] introduced a combination feature model of B-mode ultrasound images and strain elastography and showed better performance than those of models established using these two modalities alone. Tanaka et al. [55] introduced an ensemble classifier of two CNN models based on VGG19 and ResNet152 for multiple view images of one mass. Table 1 summarizes the number of cases, methods used in each paper, and the performance of the previous and proposed methods. Performance improves when deep learning is configured by combining multiple sources of ultrasound anatomical information, compared with when only one anatomical image (B-mode) is used. In addition, compared to using B-mode images alone, better performance is observed when complementary molecular information is provided, and furthermore, when patient information and the BI-RADS category are added to molecular information.
Thyroid cancer is one of the most rapidly increasing cancers. Ultrasound is used as a primary diagnostic tool for detecting and diagnosing thyroid cancer. Various features are used in ultrasound images, and as shown in Table 2, many CAD studies have been conducted recently using deep learning. Nguyen et al. [60] introduced a method of combining ResNet50-based CNN architecture and Inception-based CNN architecture with a weighted binary cross-entropy loss function. Park et al. [61] integrated seven ultrasound features (composition, echogenicity, orientation, margin, spongiform, shape, and calcification) and compared the performance with those of a support vector machine-based ultrasound CAD system and radiologists. Zhu et al. [62] proposed a deep neural network method to help radiologists differentiate Bethesda class III from Bethesda class VI, V and VI lesions. In all of these cases, the deep learning models performed better than conventional machine learning, and that the performance was better when additional features were combined.
In the field of echocardiography, studies [64] that mainly focused on distinguishing cardiac disorders are now being extended to include the detection of additional information from the cardiac view or explanations proposed regarding the grounds for the determination. Ghorbani et al. [56] used video images as network input to simultaneously analyze the data characteristics that contain spatiotemporal information and created the final output by averaging the generated output from cine frames. By analyzing local cardiac structures, enlarged left atrium, left ventricular hypertrophy, and the presence of a pacemaker lead were determined, and the positions of the area important to the determination were marked to present explanations on the output. The study also tried to estimate age, sex, weight, and height from representative views, such as the apical four-chamber view.
A chronic kidney disease (CKD) scoring system [65] using ultrasonographic parameters such as kidney length, parenchymal thickness, and echogenicity is widely used. Issues still exist, however, regarding the user’s subjective evaluation. In a study by Kuo et al. [66], a sequential configuration of two networks was presented. It was configured to average the results of 10 generated networks to predict the estimated glomerular filtration rate (GFR), a renal function index, and the features used in the prediction were linked to another network to determine CKD status. The experimental results confirmed a strong correlation between the blood creatinine-based GFR prediction and the results of the AI-based application.
Automatic determination of long head of biceps tendon inflammatory severity using ultrasound imaging was attempted by Lin et al. [67]. Input images were processed first to detect the presence of the biceps. A CNN was then used to classify the images with a detected ROI into three classes of inflammatory severity (normal and mild, moderate, or severe). It was suggested that the user’s burden can be alleviated during the determination of bicipital peritendinous effusion.
Many networks have been proposed for automatic liver fibrosis staging, with examples including a four-layer CNN with elastographic image input [68] and a METAVIR [69,70] score prediction network from B-mode images. Xue et al. [71] used a multiple modality input of B-mode and elastography images. Two networks were trained using B-mode and elastography individually, and the results of each were combined to generate fibrosis staging. It was shown that networks with multi-modal input produced better performance.
There are many deep learning-based obstetrics and gynecology applications [72]. Attempts have been made to detect abnormalities in the fetal brain. Yaqub et al. [73] reported a study where axial cross-sections of fetal brain were segmented for the craniocerebral region, and input to CNN for a two-category (normal/abnormal) classification. Suspected abnormalities were also displayed using a heat map. The traditional benign/malignant classification of ovarian cysts depended only on manually designed features [74]. In a more recent study by Zhang et al. [75], a diagnosis system to determine ovarian cysts on color Doppler ultrasound images was proposed to reduce unnecessary fine-needle aspiration evaluations. High-level features generated from a deep learning network and low-level features of texture information were combined. The experimental results indicated that the differences between malignant and benign ovarian cysts could be described by using a combination of these two feature types.
Clinical decision support solutions, traditionally referred to as CADx, have been developed gradually over the years. However, these traditional methods were not applicable as practical diagnosis tools because their existing performance generally did not satisfy doctors’ needs. Recently introduced deep learning methods are showing improved performance, enabling practical applications in clinics. Commercial AI products are being released, and their clinical validation and clinical utility are becoming increasingly important. Helping with the doctor's diagnosis is not only a form of qualitative assistance to support clinical decision-making, but also a meaningful attempt to increase workflow efficiency, as the following section explains in detail.
System workflow enhancement is relatively easy in terms of collecting training data and is less restricted regarding computational resources for real-time processing; therefore, immediate and effective applications are more readily possible than is the case for imaging, which requires real-time processing, or diagnosis, which has the burden of diagnostic accuracy and training data. AI technology incorporated in an ultrasound system is applied in the scanning and measurement/quantification processes in the system workflow, as shown in Fig. 9. Fast processing and assisted scanning all simplify the job and reduce time-consuming and repetitive tasks for medical practitioners, increase their productivity, reduce costs, and improve the efficiency of the workflow. We will introduce examples of deep learning technology applied to view recognition, scan guide, image quality assessment, and quantification and measurement at each stage of the diagnostic process of ultrasonography.
On ultrasonography, it can be difficult to determine which part of the body or organ is being scanned only with a 2D cross-sectional image. Automatic view recognition started by using a support vector machine [76] and conventional machine learning [77,78], but recently techniques that integrate deep learning have been developed and have greatly improved view recognition. For example, fully automatic classification or accurate segmentation of the left ventricle (LV) was not easy because of noise and artifacts in cardiac ultrasound images. Moreover, numerous images similar to the shape of the LV appear with a considerable variety. To recognize, segment, and track the LV in imaging sequences, a new method of integrating a faster R-CNN and an active shape model (ASM) was proposed [79]. A fast R-CNN [80], was utilized to recognize the ROI, and an ASM [81] identified the parameters that most precisely expressed the shape of the LV.
Recognizing the six standard planes in the fetal brain, which is necessary for the accurate detection of fetal brain abnormalities, has also been very difficult due to wide diversity of fetal postures, insufficient data, and similarities between the standard planes. Qu et al. [82] introduced a domain transfer learning method based on deep CNN. This framework generally outperformed those using other classical deep learning methods. In addition, the experimental results showed the effectiveness of data augmentation, especially when training data were insufficient.
Cai et al. [83] introduced an automated approach, SonoEyeNet, for the automatic recognition of standardized abdominal circumference (AC) planes on fetal ultrasonography. Built in a CNN framework, the method utilized the eye movement data of a sonographer. The movement data were collected from experienced sonographers to generate visual heat maps (visual fixation information) of each frame and used the data for identifying the correct planes. Using Sononet [84], a real-time detection technology of fetal standard scan planes in freehand ultrasound, the heatmaps and image feature maps were integrated to enhance the accuracy of AC plane detection.
Ultrasound images differ according to the user. A scan guide function is needed to assist unskilled people to take ultrasound images similarly to experienced users. Reinforcement learning is a method that maximizes the reward according to the result of an action, and has the characteristic feature of being able to reflect the user's actions and experiences in the system.
Techniques have been developed to provide a scan guide by applying reinforcement learning to an ultrasound system have been developed. Although many recent approaches have focused on developing smart ultrasound equipment that adds interpretative capabilities to existing systems, Milletari et al. [85] applied reinforcement learning to guide inexperienced users in POCUS to obtain clinically relevant images of the anatomy of interest. Jarosik and Lewandowski [86] developed a software agent that easily adapts to new conditions and informs the user on how to obtain the optimal settings of the imaging system during the scanning.
In ultrasound imaging, diagnosis is performed on standard planes. It is necessary to judge whether an image captured by the user is suitable for the standard plane. The quality of ultrasound images, for obstetric examinations as an example, is important for accurate biometric measurements. Manual quality control is a labor-intensive process that is often not practical in clinical environments. Therefore, a method that improves examination efficiency and reduces measurement errors due to inappropriate ultrasound scanning and slice selection is required.
Wu et al. [87] depicted a computerized fetal ultrasound image quality assessment (FUIQA) system to support quality management in a clinical obstetric examination. The FUIQA system was implemented with two deep CNN models, L-CNN and C-CNN. The L-CNN model located the ROI of the fetal abdomen, while the C-CNN evaluated the image quality from the goodness of depiction of the key structures of the stomach bubble and umbilical vein ROI.
In echocardiography, doctors can diagnose most heart diseases by observing the shape and movement of the heart and evaluating abnormalities in blood flow. In obstetrics, diagnostic workflows exist for fetal development measurements to estimate gestational age and to diagnose fetal growth disorders and cerebral anomalies.
Conventional measurements require manual operations with several clicks, which is a tedious, error-prone, and time-consuming job. Recently, AI-based quantification tools have been applied in a wider range of clinical applications and research is underway to achieve faster and more accurate diagnoses in combination with detection tools.
Measurements of the volume of the LV and ejection fraction (EF) in two-dimensional echocardiography have a high uncertainty due to inter-observer variability of manual measurements and acquisition errors such as apical foreshortening. Smistad et al. [88] proposed a real-time and fully automated EF measurement and foreshortening detection method. This method measured the amount of foreshortening, LV volume, and EF by employing deep learning features including view classification, cardiac cycle timing, segmentation, and landmark extraction. Furthermore, Jafari et al. [10] introduced a feasible real-time mobile application on Android mobile devices wired or wirelessly connected to a cardiac POCUS system for estimating the left ventricular ejection fraction.
Measuring the fetal growth index is a routine task, and it is important to improve the accuracy and efficiency of the work through automatic measurements [89]. Kim et al. [9] introduced a deep learning-based method for automatic evaluations of fetal head biometry by first measuring the biparietal diameter and head circumference, followed by checking plane acceptability, and finally refining the measurements. Sobhaninia et al. [90] suggested a new approach in automatic segmentation and estimation for fetal HC. Using a multi-task deep network based on the structure of Link-Net [91] and an ellipse tuner, smoother and cleaner elliptical segmentation resulted in comparison to what was obtained using a single-task network. It was recently reported that, in detecting the fetal head and abdomen, many vague images where detection seemed unlikely with traditional methods actually produced meaningful results [92], showing the potential of more robust and stable technology.
A review of the most recent applications of deep learning on ultrasound imaging applications has been presented herein. Following a brief introduction to CNNs and their domains of application, including classification, detection, segmentation, and generation, some recent studies on ultrasound imaging were summarized, focusing on the role played by deep learning in scanning, diagnosis, image enhancement, quantification and measurement, and workflow efficiency improvement. One of the most important requirements for practical use of these technologies in ultrasonography is real-time implementation. The availability of peripheral computational processing technology, therefore, is a key ingredient for rapid adaptation and usage.
Deep learning-based diagnosis undoubtedly has tremendous future potential. It will surely expand and provide doctors, and society as a whole, with various benefits including better accuracy, efficient performance, and cost reduction. However, some hurdles should be overcome. Insufficient accumulation of medical imaging data could cause difficulties in verifying clinical validity and utility for practical purposes [93]. For the same reason, but from a different point of view, regulatory agencies such as the Food and Drug Administration (FDA), China National Medical Products Administration (NMPA), and the South Korean Ministry of Food and Drug Safety (MFDS) are working on risk management and discussing whether deep learning based algorithms should be allowed to be incorporated into medical devices. There is also a longstanding controversy regarding the proper level of accuracy in AI diagnoses. A shared understanding now exists that AI can, even if not at the level of an expert, still reduce simple human errors and contribute to enhancing average diagnostic accuracy by providing a second opinion to a doctor’s decision [94]. Furthermore, the new development of multi-parameter and multi-modal diagnoses may possibly lead to the next level of comprehensive diagnostic tools for medical professionals.
Image quality enhancement due to deep learning is expected to start with postprocessing of the images first and eventually to cover ultrasound beamforming, contributing to fundamental image quality improvement. The application of advanced beamforming technology, which has been studied for several decades but has not been successfully applied in general, could also be expected through deep learning. Workflow enhancement is the most active domain of applications, especially for commercial implementation. Recently, regulatory agencies, including the FDA and MFDS, have been cautiously easing regulations on CADe. These changes would simplify regulatory review and give patients more timely access to CADe software applications. The FDA believes that these special controls will provide a reasonable assurance of safety and effectiveness [95]. Easing regulations in this field implies that qualified doctors can enhance their workflow and improve productivity by routinely using these technologies in their daily practice. Improved productivity will be perceived by not only healthcare professionals, but by society as a whole in the form of cost reduction and financial efficiency.
Finally it should be mentioned that government and healthcare authorities will play the paramount role in these innovations. Standardized and unified guidelines and regulations have yet to be developed. Active discussions and workshops are going on among many parties involved, such as the International Medical Device Regulators Forum. Participation in such initiatives is strongly recommended for academia, industry, research centers, and governing institutions.
NotesAuthor Contributions Conceptualization: Bang WC, Yi J, Kim DW. Data acquisition: Park MH, Seong YK, Kim KS, Kwon JH, Lee J, Kang HK. Data analysis or interpretation: Park MH, Seong YK, Kim KS, Kwon JH, Lee J, Kang HK. Drafting of the manuscript: Park MH, Seong YK, Kim KS, Kwon JH, Lee J, Kang HK, Yi J, Ha K, Ahn B. Critical revision of the manuscript: Bang WC, Yi J, Hah Z, Kim DW. Approval of the final version of the manuscript: Bang WC. Conflict of InterestAll the authors are employees of Samsung Electronics Co., Ltd., or Samsung Medison Co., Ltd. References1. Carneiro G, Nascimento JC. Combining multiple dynamic models and deep learning architectures for tracking the left ventricle endocardium in ultrasound data. IEEE Trans Pattern Anal Mach Intell 2013;35:2592–2607.
2. Carneiro G, Nascimento JC, Freitas A. The segmentation of the left ventricle of the heart from ultrasound data using deep learning architectures and derivative-based search methods. IEEE Trans Image Process 2012;21:968–982.
3. Solomon O, Cohen R, Zhang Y, Yang Y, He Q, Luo J, et al. Deep unfolded robust PCA with application to clutter suppression in ultrasound. IEEE Trans Med Imaging 2020;39:1051–1063.
4. Yoon YH, Khan S, Huh J, Ye JC. Efficient B-mode ultrasound image reconstruction from sub-sampled RF data using deep learning. IEEE Trans Med Imaging 2019;38:325–336.
5. Prevost R, Salehi M, Jagoda S, Kumar N, Sprung J, Ladikos A, et al. 3D freehand ultrasound without external tracking using deep learning. Med Image Anal 2018;48:187–202.
6. Feigin M, Freedman D, Anthony BW. A deep learning framework for single-sided sound speed inversion in medical ultrasound. IEEE Trans Biomed Eng 2020;67:1142–1151.
7. Luchies AC, Byram BC. Training improvements for ultrasound beamforming with deep neural networks. Phys Med Biol 2019;64:045018.
8. Yu H, Ding M, Zhang X, Wu J. PCANet based nonlocal means method for speckle noise removal in ultrasound images. PLoS One 2018;13:e0205390.
9. Kim HP, Lee SM, Kwon JY, Park Y, Kim KC, Seo JK. Automatic evaluation of fetal head biometry from ultrasound images using machine learning. Physiol Meas 2019;40:065009.
10. Jafari MH, Girgis H, Van Woudenberg N, Liao Z, Rohling R, Gin K, et al. Automatic biplane left ventricular ejection fraction estimation with mobile point-of-care ultrasound using multi-task learning and adversarial training. Int J Comput Assist Radiol Surg 2019;14:1027–1037.
11. Buda M, Wildman-Tobriner B, Castor K, Hoang JK, Mazurowski MA. Deep learning-based segmentation of nodules in thyroid ultrasound: improving performance by utilizing markers present in the images. Ultrasound Med Biol 2020;46:415–421.
12. Loram I, Siddique A, Sanchez MB, Harding P, Silverdale M, Kobylecki C, et al. Objective analysis of neck muscle boundaries for cervical dystonia using ultrasound imaging and deep learning. IEEE J Biomed Health Inform 2020;24:1016–1027.
13. Park H, Lee HJ, Kim HG, Ro YM, Shin D, Lee SR, et al. Endometrium segmentation on transvaginal ultrasound image using key-point discriminator. Med Phys 2019;46:3974–3984.
14. Ryou H, Yaqub M, Cavallaro A, Papageorghiou AT, Alison Noble J. Automated 3D ultrasound image analysis for first trimester assessment of fetal health. Phys Med Biol 2019;64:185010.
15. Yap MH, Pons G, Marti J, Ganau S, Sentis M, Zwiggelaar R, et al. Automated breast ultrasound lesions detection using convolutional neural networks. IEEE J Biomed Health Inform 2018;22:1218–1226.
16. Yap MH, Goyal M, Osman FM, Marti R, Denton E, Juette A, et al. Breast ultrasound lesions recognition: end-to-end deep learning approaches. J Med Imaging (Bellingham) 2019;6:011007.
17. Yin S, Peng Q, Li H, Zhang Z, You X, Liu H, et al. Multi-instance deep learning with graph convolutional neural networks for diagnosis of kidney diseases using ultrasound imaging. In: Greenspan H, Tanno R, Erdt M, eds. Uncertainty for safe utilization of machine learning in medical imaging and clinical image-based procedures. Lecture notes in computer science, No. 11840. Cham: Springer, 2019:146–154.
18. Lei Y, Tian S, He X, Wang T, Wang B, Patel P, et al. Ultrasound prostate segmentation based on multidirectional deeply supervised V-Net. Med Phys 2019;46:3194–3206.
20. Akkus Z, Cai J, Boonrod A, Zeinoddini A, Weston AD, Philbrick KA, et al. A survey of deep-learning applications in ultrasound: artificial intelligence-powered ultrasound for improving clinical workflow. J Am Coll Radiol 2019;16(9 Pt B):1318–1328.
22. Caffe: a deep learning framework [Internet]. Berkeley, CA: University of California, 2020. [cited 2020 Aug 30]. Available from: http://caffe.berkeleyvision.org/.
23. TensorFlow: TensorFlow is an end-to-end open source platform for machine learning [Internet]. Mountain View, CA: Google, 2020. [cited 2020 Aug 30]. Available from:
https://www.tensorflow.org/.
24. Keras: the high-level API of TensorFlow 2.0 [Internet]. San Francisco, CA: GitHub, 2020. [cited 2020 Aug 30]. Available from: https://keras.io/.
25. Torch: a scientific computing framework [Internet]. San Francisco, CA: GitHub, 2017. [cited 2020 Aug 30]. Available from: http://torch.ch/.
26. MXNet: A deep learning framework designed for both efficiency and flexibility [Internet]. San Francisco, CA: GitHub, 2020. [cited 2020 Aug 30]. Available from: https://mxnet.apache.org/.
27. Krizhevsky A, Sutskever I, Hinton GE. ImageNet classification with deep convolutional neural networks. Adv Neural Inf Process Syst 2012;25:1097–1105.
28. Simonyan K, Zisserman A. Very deep convolutional networks for large-scale image recognition. Preprint at https://arxiv.org/abs/1409.1556 (2015).
29. Szegedy C, Liu W, Jia Y, Sermanet P, Reed S, Anguelov D, et al. Going deeper with convolutions. Proc IEEE Comput Soc Conf Comput Vis Pattern Recognit 2015;2015:1–9.
30. He K, Zhang X, Ren S, Sun J. Deep residual learning for image recognition. Proc IEEE Comput Soc Conf Comput Vis Pattern Recognit 2016;2016:770–778.
31. Huang G, Liu Z, van der Maaten L, Weinberger KQ. Densely connected convolutional networks. Proc IEEE Comput Soc Conf Comput Vis Pattern Recognit 2017;2017:4700–4708.
32. Girshick R, Donahue J, Darrell T, Malik J. Rich feature hierarchies for accurate object detection and semantic segmentation. Proc IEEE Comput Soc Conf Comput Vis Pattern Recognit 2014;2014:580–587.
33. Redmon J, Farhadi A. YOLO9000: better, faster, stronger. Proc IEEE Comput Soc Conf Comput Vis Pattern Recognit 2017;2017:7263–7271.
34. Redmon J, Farhadi A. YOLOv3: an incremental improvement. Preprint at https://arxiv.org/abs/1804.02767 (2018).
35. Liu W, Anguelov A, Erhan D, Szegedy C, Reed S, Fu C, et al. SSD: single shot multibox detector. In: Leibe B, Matas J, Sebe N, Welling M, eds. Computer Vision - ECCV 2016. Lecture notes in computer science, Vol. 9905. Cham: Springer, 2016:12–37.
36. Ronneberger O, Fischer P, Brox T. U-net: convolutional networks for biomedical image segmentation. Med Image Comput Comput Assist Interv 2015;9351:234–241.
37. Long J, Shelhamer E, Darrell T. Fully convolutional networks for semantic segmentation. Proc IEEE Comput Soc Conf Comput Vis Pattern Recognit 2015;2015:3431–3440.
38. Noh H, Hong S, Han B. Learning deconvolution network for semantic segmentation. IEEE Int Conf Comput Vis Workshops 2015;2015:1520–1528.
39. Long J, Shelhamer E, Darrell T. Fully convolutional networks for semantic segmentation. Proc IEEE Comput Soc Conf Comput Vis Pattern Recognit 2015;2015:3431–3440.
40. Kingma DP, Welling M. Auto-encoding variational bayes. Preprint at https://arxiv.org/abs/1312.6114 (2014).
41. Vincent P, Larochelle H, Lajoie I, Bengio Y, Manzagol PA. Stacked denoising autoencoders: learning useful representations in a deep network with a local denoising criterion. J Mach Learn Res 2010;11:3371–3408.
42. Radford A, Metz L, Chintala S. Unsupervised representation learning with deep convolutional generative adversarial networks. Preprint at https://arxiv.org/abs/1511.06434 (2016).
43. Ledig C, Theis L, Huszar F, Caballero J, Cunningham A, Acosta A, et al. Photo-realistic single image super-resolution using a generative adversarial network. Proc IEEE Comput Soc Conf Comput Vis Pattern Recognit 2017;2017:4681–4690.
44. Rawat W, Wang Z. Deep convolutional neural networks for image classification: a comprehensive review. Neural Comput 2017;29:2352–2449.
45. Synnevag JF, Austeng A, Holm S. Adaptive beamforming applied to medical ultrasound imaging. IEEE Trans Ultrason Ferroelectr Freq Control 2007;54:1606–1613.
46. Luijten B, Cohen R, de Bruijn FJ, Schmeitz HA, Mischi M, Eldar YC, et al. Deep learning for fast adaptive beamforming. Proc IEEE Int Conf Acoust Speech Signal Process 2019;2019:1333–1337.
47. Liu X, Zhou T, Lu M, Yang Y, He Q, Luo J. Deep Learning for Ultrasound Localization Microscopy. IEEE Trans Med Imaging 2020;39:3064–3078.
48. Kervrann C, Boulanger J, Coupe P. Bayesian non-local means filter, image redundancy and adaptive dictionaries for noise removal. In: Sgallari F, Murli A, Paragios N, eds. Scale Space and Variational Methods in Computer Vision. Lecture notes in computer science, Vol. 4485. Berlin: Springer, 2007:520–532.
49. Khan S, Huh J, Ye JC. Adaptive and compressive beamforming using deep learning for medical ultrasound. IEEE Trans Ultrason Ferroelectr Freq Control 2020;67:1558–1572.
50. Du B, Wang J, Zheng H, Xiao C, Fang S, Lu M, et al. A novel transcranial ultrasound imaging method with diverging wave transmission and deep learning approach. Comput Methods Programs Biomed 2020;186:105308.
51. Huang O, Long W, Bottenus N, Lerendegui M, Trahey GE, Farsiu S, et al. MimickNet, Mimicking clinical image post-processing under black-box constraints. IEEE Trans Med Imaging 2020;39:2277–2286.
52. Jafari MH, Girgis H, Van Woudenberg N, Moulson N, Luong C, Fung A, et al. Cardiac point-of-care to cart-based ultrasound translation using constrained CycleGAN. Int J Comput Assist Radiol Surg 2020;15:877–886.
53. Wildeboer RR, Van Sloun RJ, Mannaerts CK, Moraes PH, Salomon G, Chammas MC, et al. Synthetic elastography using B-mode ultrasound through a deep fully-convolutional neural network. IEEE Trans Ultrason Ferroelectr Freq Control 2020 Mar 24 [Epub]. https://doi.org/10.1109/TUFFC.2020.2983099.
54. Liao WX, He P, Hao J, Wang XY, Yang RL, An D, et al. Automatic identification of breast ultrasound image based on supervised block-based region segmentation algorithm and features combination migration deep learning model. IEEE J Biomed Health Inform 2020;24:984–993.
55. Tanaka H, Chiu SW, Watanabe T, Kaoku S, Yamaguchi T. Computer-aided diagnosis system for breast ultrasound images using deep learning. Phys Med Biol 2019;64:235013.
56. Ghorbani A, Ouyang D, Abid A, He B, Chen JH, Harrington RA, et al. Deep learning interpretation of echocardiograms. NPJ Digit Med 2020;3:10.
57. Ye JC, Han Y, Cha E. Deep convolutional framelets: a general deep learning framework for inverse problems. SIAM J Imaging Sci 2018;11:991–1048.
58. Zheng X, Yao Z, Huang Y, Yu Y, Wang Y, Liu Y, et al. Deep learning radiomics can predict axillary lymph node status in early-stage breast cancer. Nat Commun 2020;11:1236.
59. Sun Q, Lin X, Zhao Y, Li L, Yan K, Liang D, et al. Deep learning vs. radiomics for predicting axillary lymph node metastasis of breast cancer using ultrasound images: don't forget the peritumoral region. Front Oncol 2020;10–53.
60. Nguyen DT, Kang JK, Pham TD, Batchuluun G, Park KR. Ultrasound image-based diagnosis of malignant thyroid nodule using artificial intelligence. Sensors (Basel) 2020;20:1822.
61. Park VY, Han K, Seong YK, Park MH, Kim EK, Moon HJ, et al. Diagnosis of thyroid nodules: performance of a deep learning convolutional neural network model vs. radiologists. Sci Rep 2019;9:17843.
62. Zhu Y, Sang Q, Jia S, Wang Y, Deyer T. Deep neural networks could differentiate Bethesda class III versus class IV/V/VI. Ann Transl Med 2019;7:231.
63. Nguyen DT, Pham TD, Batchuluun G, Yoon HS, Park KR. Artificial intelligence-based thyroid nodule classification using information from spatial and frequency domains. J Clin Med 2019;8:1976.
64. Zhang J, Gajjala S, Agrawal P, Tison GH, Hallock LA, Beussink-Nelson L, et al. Fully automated echocardiogram interpretation in clinical practice. Circulation 2018;138:1623–1635.
65. Yaprak M, Cakir O, Turan MN, Dayanan R, Akin S, Degirmen E, et al. Role of ultrasonographic chronic kidney disease score in the assessment of chronic kidney disease. Int Urol Nephrol 2017;49:123–131.
66. Kuo CC, Chang CM, Liu KT, Lin WK, Chiang HY, Chung CW, et al. Automation of the kidney function prediction and classification through ultrasound-based kidney imaging using deep learning. NPJ Digit Med 2019;2:29.
67. Lin BS, Chen JL, Tu YH, Shih YX, Lin YC, Chi WL, et al. Using deep learning in ultrasound imaging of bicipital peritendinous effusion to grade inflammation severity. IEEE J Biomed Health Inform 2020;24:1037–1045.
68. Wang K, Lu X, Zhou H, Gao Y, Zheng J, Tong M, et al. Deep learning radiomics of shear wave elastography significantly improved diagnostic performance for assessing liver fibrosis in chronic hepatitis B: a prospective multicentre study. Gut 2019;68:729–741.
69. Castera L, Vergniol J, Foucher J, Le Bail B, Chanteloup E, Haaser M, et al. Prospective comparison of transient elastography, Fibrotest, APRI, and liver biopsy for the assessment of fibrosis in chronic hepatitis C. Gastroenterology 2005;128:343–350.
70. Rousselet MC, Michalak S, Dupre F, Croue A, Bedossa P, Saint-Andre JP, et al. Sources of variability in histological scoring of chronic viral hepatitis. Hepatology 2005;41:257–264.
71. Xue LY, Jiang ZY, Fu TT, Wang QM, Zhu YL, Dai M, et al. Transfer learning radiomics based on multimodal ultrasound imaging for staging liver fibrosis. Eur Radiol 2020;30:2973–2983.
72. Iftikhar P, Kuijpers MV, Khayyat A, Iftikhar A, DeGouvia De Sa M. Artificial intelligence: a new paradigm in obstetrics and gynecology research and clinical practice. Cureus 2020;12:e7124.
73. Yaqub M, Kelly B, Papageorghiou AT, Noble JA. A deep learning solution for automatic fetal neurosonographic diagnostic plane verification using clinical standard constraints. Ultrasound Med Biol 2017;43:2925–2933.
74. Khazendar S, Sayasneh A, Al-Assam H, Du H, Kaijser J, Ferrara L, et al. Automated characterisation of ultrasound images of ovarian tumours: the diagnostic accuracy of a support vector machine and image processing with a local binary pattern operator. Facts Views Vis Obgyn 2015;7:7–15.
75. Zhang L, Huang J, Liu L. Improved deep learning network based in combination with cost-sensitive learning for early detection of ovarian cancer in color ultrasound detecting system. J Med Syst 2019;43:251.
76. Chang RF, Wu WJ, Moon WK, Chen DR. Improvement in breast tumor discrimination by support vector machines and speckle-emphasis texture analysis. Ultrasound Med Biol 2003;29:679–686.
77. Attia MW, Abou-Chadi FE, Moustafa HE, Mekky N. Classification of ultrasound kidney images using PCA and neural networks. Int J Adv Comput Sci Appl 2015;6:53–57.
78. Bridge CP, Ioannou C, Noble JA. Automated annotation and quantitative description of ultrasound videos of the fetal heart. Med Image Anal 2017;36:147–161.
79. Hsu WY. Automatic left ventricle recognition, segmentation and tracking in cardiac ultrasound image sequences. IEEE Access 2019;7:140524–140533.
81. Cootes TF, Taylor CJ, Cooper DH, Graham J. Active shape models: their training and application. Comput Vis Image Underst 1995;61:38–59.
82. Qu R, Xu G, Ding C, Jia W, Sun M. Deep learning-based methodology for recognition of fetal brain standard scan planes in 2D ultrasound images. IEEE Access 2019;8:44443–44451.
83. Cai Y, Sharma H, Chatelain P, Noble JA. SonoEyeNet: standardized fetal ultrasound plane detection informed by eye tracking. Proc IEEE Int Symp Biomed Imaging 2018;2018:1475–1478.
84. Baumgartner CF, Kamnitsas K, Matthew J, Fletcher TP, Smith S, Koch LM, et al. SonoNet: real-time detection and localisation of fetal standard scan planes in freehand ultrasound. IEEE Trans Med Imaging 2017;36:2204–2215.
85. Milletari F, Birodkar V, Sofka M. Straight to the point: reinforcement learning for user guidance in ultrasound. In: Wang Q, Gomez A, Hutter J, McLeod K, Zimmer V, Zettinig O, eds. Smart ultrasound imaging and perinatal, preterm and paediatric image analysis. Cham: Springer, 2019:3–10.
86. Jarosik P, Lewandowski M. Automatic ultrasound guidance based on deep reinforcement learning. IEEE Int Ultrason Symp 2019;2019:475–478.
87. Wu L, Cheng JZ, Li S, Lei B, Wang T, Ni D. FUIQA: fetal ultrasound image quality assessment with deep convolutional networks. IEEE Trans Cybern 2017;47:1336–1349.
88. Smistad E, Ostvik A, Salte IM, Melichova D, Nguyen TM, Haugaa K, et al. Real-time automatic ejection fraction and foreshortening detection using deep learning. IEEE Trans Ultrason Ferroelectr Freq Control 2020 Mar 16 [Epub]. https://doi.org/10.1109/TUFFC.2020.2981037.
89. Espinoza J, Good S, Russell E, Lee W. Does the use of automated fetal biometry improve clinical work flow efficiency? J Ultrasound Med 2013;32:847–850.
90. Sobhaninia Z, Rafiei S, Emami A, Karimi N, Najarian K, Samavi S, et al. Fetal ultrasound image segmentation for measuring biometric parameters using multi-task deep learning. Annu Int Conf IEEE Eng Med Biol Soc 2019;2019:6545–6548.
91. Chaurasia A, Culurciello E. LinkNet: exploiting encoder representations for efficient semantic segmentation. 2017 IEEE Visual Communications and Image Processing (VCIP); 2017 Dec 10-13; St. Petersburg, FL, USA. New York: Institute of Electrical and Electronics Engineers, 2017. 518–521.
92. Kwon JY. Feasibility of improved algorithm-based BiometryAssist in fetal biometric measurement (white paper) [Internet]. Seongnam: Samsung Healthcare, 2019. [cited 2020 Aug 30]. Available from: https://www.samsunghealthcare.com/en/knowledge_hub/clinical_library/white_paper.
93. Kim DW, Jang HY, Kim KW, Shin Y, Park SH. Design characteristics of studies reporting the performance of artificial intelligence algorithms for diagnostic analysis of medical images: results from recently published papers. Korean J Radiol 2019;20:405–410.
94. Allen B Jr, Seltzer SE, Langlotz CP, Dreyer KP, Summers RM, Petrick N, et al. A road map for translational research on artificial intelligence in medical imaging: from the 2018 National Institutes of Health/RSNA/ACR/The Academy Workshop. J Am Coll Radiol 2019;16(9 Pt A):1179–1189.
95. Ridley E. FDA finalizes easier rules for CADe software [Internet]. Arlington, VA: AuntMinnie, 2020. [cited 2020 Aug 30]. Available from: https://www.auntminnie.com/index.aspx?sec=sup&sub=aic&pag=dis&ItemID=127921.
The number of published articles with "deep learning" in PubMed (A) and both "deep learning" and "ultrasound" (B).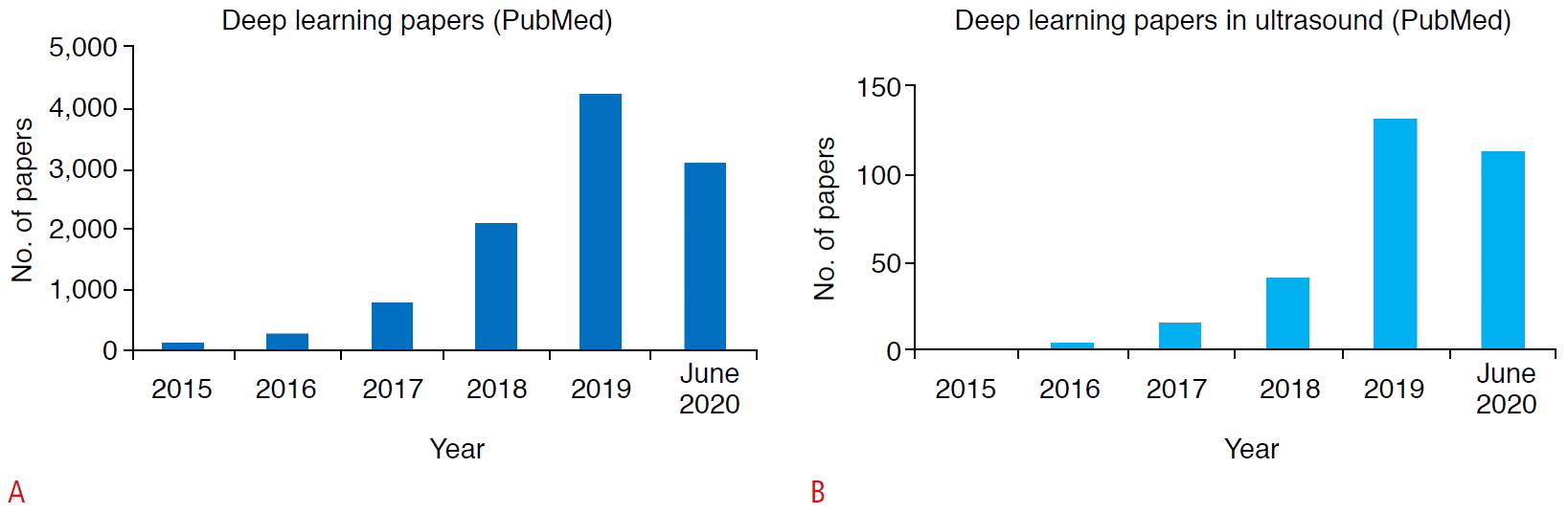 Fig. 1.Basic components of a convolutional neural network.It consists of convolutional layers performing filtering inputs (A) and pooling layers for down-sampling images (B). Through the training process, the optimal parameters of convolutional weights can be determined.
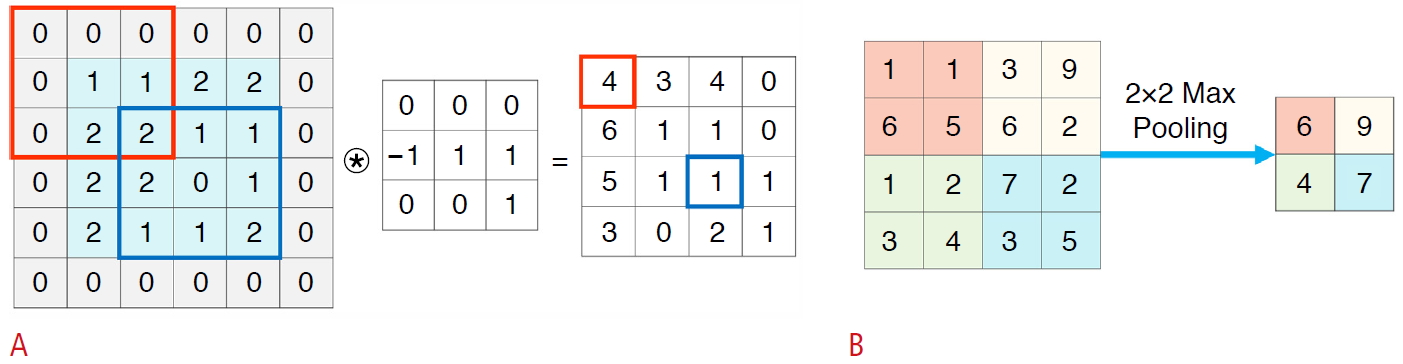 Fig. 2.Architecture of U-Net for segmentation.U-Net consists of pairs of encoder (compressing the data) and decoder (uncompressing the data). Detailed segmentation is possible by connecting the encoder's scale-specific features to the decoder's features.
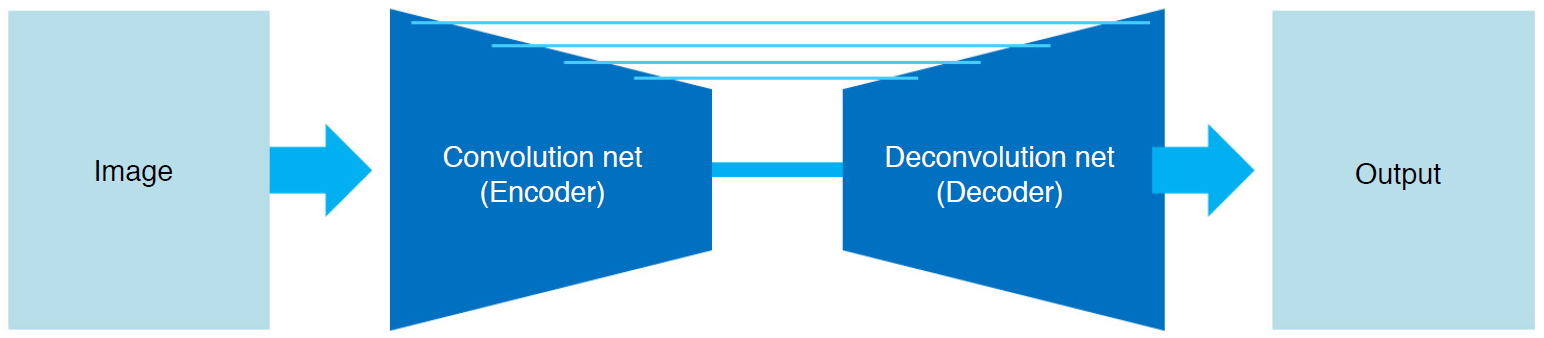 Fig. 5.Basic architecture of a generative adversarial network.It consists of a generator network that creates an unseen image from an input random variable and a discriminator network that distinguishes whether the image is real or created.
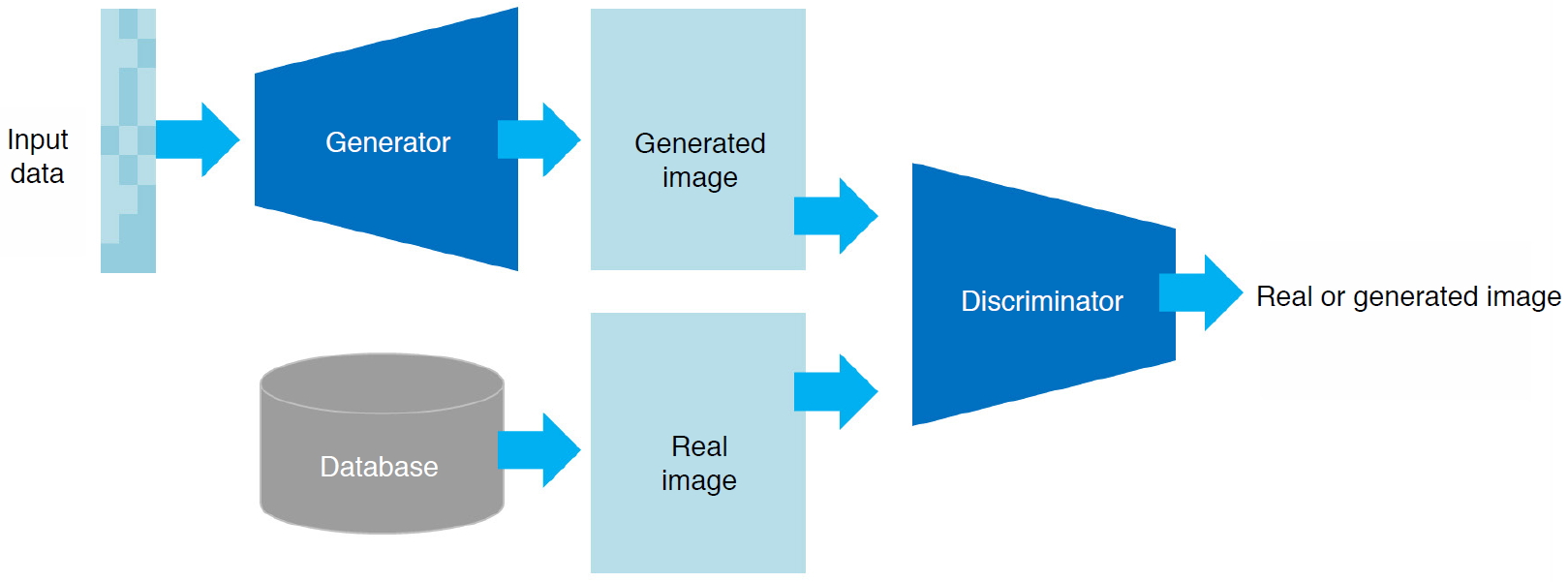 Fig. 6.Examples of classification, detection, segmentation, and generation applied to ultrasound images.Classification of malignancy of breast lesion (A), detection and segmentation of the ulnar nerve (UN) (B), circumference segmentation of the fetal abdomen (C), and examples of generated ultrasound images resembling breast lesions (D) are shown.
 Fig. 7.Transfer learning approach.A pretrained model based on a large dataset can be effectively used for applications with a smaller dataset.
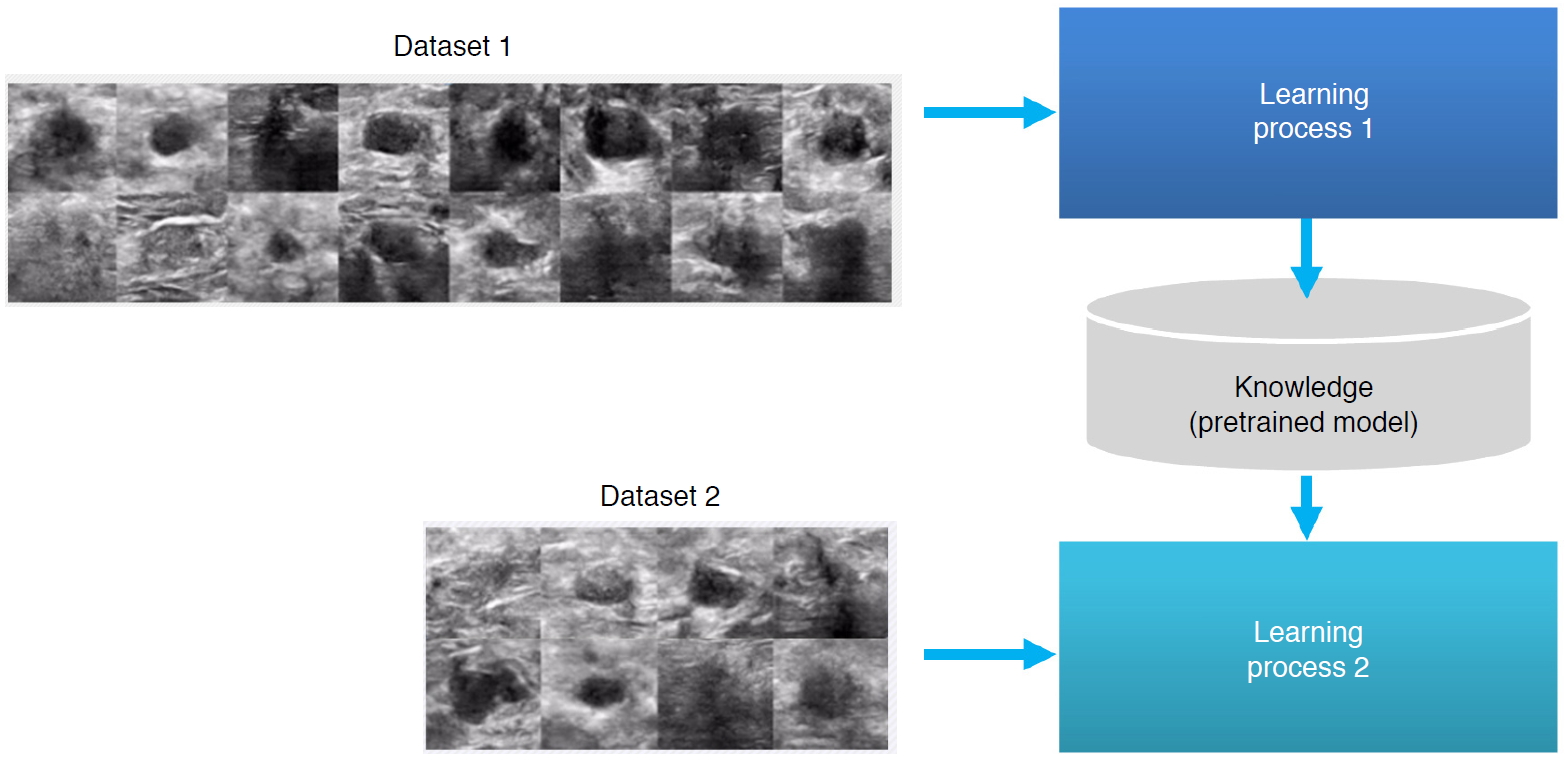 Fig. 8.A simplified diagram of the diagnostic workflow in ultrasonography when deep learning is involved.CADe, computer-aided detection; CADq, computer-aided quantification; CADx, computer-aided diagnosis; CADt, computer-aided triage.
 Fig. 9.Table 1.Deep learning research on breast diagnosis
Table 2.Deep learning research on thyroid diagnosis
|



 Print
Print facebook
facebook twitter
twitter Linkedin
Linkedin google+
google+

 Download Citation
Download Citation PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC




