AbstractPurposeThis study aimed to identify useful shear wave elastography (SWE) parameters for differentiating breast cancer and predicting associated immunohistochemical factors and subtypes.
MethodsFrom November 2018 to February 2019, a total of 211 breast lesions from 190 patients who underwent conventional breast ultrasonography and SWE were included. The Breast Imaging Reporting and Data System categories and qualitative and quantitative SWE parameters for each lesion were obtained. Pathologic results including immunohistochemical factors were evaluated. The diagnostic performance of each parameter and its correlation with histological characteristics, immunohistochemical factors, and subtypes of breast cancer were analyzed using analysis of variance, the independent t test, the Fisher exact test, logistic regression analysis, and the DeLong method.
ResultsAmong 211 breast lesions, 82 were malignant, and 129 were benign. Of the SWE parameters, Emax showed the highest area under the curve (AUC) for differentiating malignant from benign lesions (AUC, 0.891; cut-off>50.85). Poor tumor differentiation and progesterone receptor-negativity were correlated with higher SDmean and Emax (P<0.05). Ki-67-positive breast cancer showed higher SDmean and a heterogeneous color distribution (P<0.05). Ki-67 and cytokeratin 5/6-positive breast cancers showed higher Emax/Efat ratios (P<0.05). Luminal B, human epidermal growth factor receptor 2-enriched, and triple-negative (non-basal) subtypes showed somewhat higher SDmean values than the luminal A and triple-negative (basal) subtypes (P=0.028).
Breast cancer is a heterogeneous disease with various immunohistochemical subtypes. Immunohistochemical factors can affect the clinical course, treatment response, and prognosis of breast cancer patients [1]. Estrogen receptor (ER) and progesterone receptor (PR) are useful markers when selecting patients for hormone therapy. Human epidermal growth factor receptor 2 (HER2) is a valuable immunohistochemical predictor of response to trastuzumab [2]. Ki-67 is another immunohistochemical factor associated with cell proliferation, which is known to be a poor prognostic factor. Several studies have revealed that measuring Ki-67-positive cells during treatment can improve predictions of treatment efficacy in both antihormonal therapy and chemotherapy [3,4]. In addition, the pre-treatment Ki-67 index has been found to be associated with the survival rate after neoadjuvant chemotherapy and with the response to neoadjuvant chemotherapy [5,6].
Breast elastography has been used to evaluate breast lesions by tissue elasticity with high sensitivity and specificity [7]. Shear wave elastography (SWE) is a quantitative technique that can calculate tissue elasticity by measuring the velocity of shear waves propagating in the tissue [8]. Unlike strain elastography, SWE can quantitatively and reproducibly measure tissue stiffness without examiner-dependence [9,10]. Therefore, SWE can produce more objective measurements, making it more useful for determining tissue characteristics [11]. Some studies have shown that breast cancers tend to be stiffer on SWE, which can be used in differentiating malignant from benign lesions [12].
Recently, associations between tumor stiffness and immunohistochemical profiles have been investigated using SWE. Ganau et al. [13] reported that there appeared to be a trend toward lower Emax and Emean values in tumors with aggressive phenotypes such as HER2 overexpression. In contrast, Youk et al. [14] found no significant correlations between mean elasticity values and the immunohistochemical profile. Chang et al. [15] found that tumors with HER2-positive and triple-negative immunohistochemical profiles were stiffer than ER-positive subtypes. Denis et al. [16] also reported that the Ki-67 proliferation index and histological grade were factors that influenced mean elasticity when differentiating between luminal subtypes.
Building upon these previous studies, we analyzed various SWE parameters in breast lesions. The objective of this study was to identify useful SWE parameters for differentiating breast cancer and predicting its immunohistochemical factors and subtypes.
This study was approved by our Institutional Review Board, and the requirement for informed consent was waived. This retrospective study included patients with breast lesions of Breast Imaging Reporting and Data System (BI-RADS) category 3 or above that were scheduled for biopsy, who underwent conventional breast ultrasonography (US) and SWE from November 2018 to February 2019. Initially, 237 breast lesions from 216 patients were included as subjects of this study. However, 26 lesions from 26 patients were excluded for the following reasons: failure to obtain proper SWE images (skin or superficial lesion, n=4; subareolar lesion, n=3; bulging mass larger than 2 cm, n=5; and axillary lesion, n=2) or absence of pathologic results (n=12). In total, 211 lesions from 190 patients were ultimately enrolled. All malignant lesions were surgically resected and benign lesions were biopsied or surgically resected.
Breast lesions were examined using a Toshiba Aplio i700 device (Toshiba Medical Systems Europe BV, Zoetermeer, The Netherlands) equipped with a PLT-705BT probe (a linear probe with a frequency range of 3-11 MHz). Examinations were performed by three expert radiologists specializing in breast imaging, each with at least 7 years of experience in breast US and a 2-month learning period for consensus in this SWE study. Consecutive breast lesions were evaluated by gray-scale US for BI-RADS categorization. SWE was consecutively performed on lesions with a BI-RADS category of over 3, and qualitative and quantitative parameters were assessed. Images of two orthogonal planes (axial and sagittal planes) were obtained and a representative image with minimal anisotropy artifacts was selected. The color distribution pattern of the lesion, one of the qualitative parameters of SWE, was determined according to the Tozaki classification [17], which contains four patterns: pattern 1 (coded blue homogeneously), pattern 2 (vertical stripe pattern artifacts), pattern 3 (a localized colored area at the margin of the lesion), and pattern 4 (heterogeneously colored areas in the interior of the lesion). Patterns 3 and 4 are defined as a heterogeneous color distribution. The quantitative parameters of SWE analyzed in this study included the mean elasticity value (Emean) and its standard deviation (SDmean), the maximum elasticity value (Emax) and its standard deviation (SDmax), and the elasticity ratio between the lesion and the adjacent fat lobule (Emean/Efat ratio and Emax/Efat ratio). Emean was obtained by delicately drawing the region of interest (ROI) along the margin of the entire mass and then calculating the mean elasticity value of the lesion (Figs. 1, 2). The ROI was drawn along the margin of the lesion, which is the edge of the hypoechoic area in a hypoechoic mass, so that it did not focus or necessarily include the stiff rim area outside the lesion. Emax was obtained by placing a small (2-3 mm) round ROI on the hardest region of the mass as determined by a visual assessment (Figs. 1, 2).
Pathologic reports of biopsy or surgical specimens were reviewed to determine tumor size, depth, histologic type and grade, lymph node metastasis, and immunohistochemical subtypes. The following biomarkers were evaluated as immunohistochemical factors: ER, PR, HER2, Ki-67, epidermal growth factor receptor (EGFR), and cytokeratin (CK) 5/6. ER, PR, HER2, Ki-67, and EGFR immunohistochemical staining was performed on an automated Ventana BenchmarkXT slide stainer (Ventana, Tucson, AZ, USA), using primary antibodies against ER (prediluted, SP1, Ventana), PR (prediluted, 1E2, Ventana), HER2 (prediluted, 4B5, Ventana), Ki-67 (prediluted, MIB-1, Ventana), and EGFR (prediluted; 3C6, Ventana). CK5/6 (1:200, D5/16 B4, Dako, Carpinteria, CA, USA) immunohistochemical staining was performed on a Dako Omnis device (Dako). A cut-off value of Ōēź1% was used to define ER and PR positivity. The intensity of HER2 expression was scored semiquantitatively as 0, 1, 2, or 3. Tumors with a score of 3 were classified as HER2-positive and those with a score of 0 or 1 were classified as HER2-negative. Gene amplification was used to determine HER2 status for tumors with a score of 2 [18]. Positive Ki-67 expression was defined as Ki-67 positivity in Ōēź14% of cancer cell nuclei. A cut-off value of Ōēź1% was used to define EGFR and CK5/6 positivity. The immunohistochemical subtypes were categorized into the following types according to the 2013 St. Gallen International Breast Cancer Conferences [19]: luminal A (ER or PR+, HER2-, and Ki-67 low), luminal B (ER or PR+, HER2+, and/or Ki-67 high), HER2+ (ER-, PR-, and HER2+), triple-negative basal-like (ER-, PR-, HER2-, and EGFR or CK5/6+), and triple-negative non-basal like (ER-, PR-, HER2-, EGFR-, and CK5/6-).
All statistical analyses were performed using SAS version 9.4 (SAS Institute, Cary, NC, USA). A P-value below 0.05 was considered to indicate statistical significance. The SWE parameters were compared between malignant and benign tumors using analysis of variance (ANOVA), the independent t test, or the Fisher exact test. Immunohistochemical factors and subtypes of breast cancer were correlated with SWE parameters using ANOVA, the Fisher exact test, and logistic regression analysis. The area under the receiver operating characteristic curve (AUC) using logistic regression analysis was plotted to assess the cut-off value for each SWE parameter. Diagnostic performance was calculated by obtaining AUC, sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV). Associations between malignancy and baseline variables were determined by logistic regression analysis. The DeLong method was used to analyze the AUC of SWE parameters according to category classification, depth, and size [20].
The baseline characteristics of the patients and lesions of interest are summarized in Table 1. The mean age was 48.0┬▒11.2 years (range, 20 to 81 years). Of the 211 breast lesions, 82 were malignant, including 66 invasive ductal cancers, 13 ductal carcinomas in situ, two invasive lobular carcinomas, and one malignant phyllodes tumor. The other 129 were benign lesions, including fibroadenoma (n=50), fibrocystic change (n=47), intraductal papilloma (n=10), sclerosing adenosis (n=4), chronic inflammation (n=4), benign phyllodes tumor (n=3), hamartoma (n=2), atypical ductal hyperplasia (n=2), stromal fibrosis (n=2), tubular adenoma (n=1), fat necrosis (n=1), columnar cell hyperplasia (n=1), usual ductal hyperplasia (n=1), and mucocele-like lesion (n=1). The mean size was 18.3┬▒8.6 mm for the 82 malignant lesions and 12.4┬▒6.8 mm (P<0.001) for the 129 benign lesions. The mean depth was 14.0┬▒3.8 mm for the malignant lesions and 12.0┬▒4.5 mm (P<0.001) for the benign lesions. None of the BI-RADS category 3 lesions were confirmed as malignant, and all 82 malignant lesions were BI-RADS category 4A or above. With a cut-off value of Ōēźcategory 4A, the sensitivity, specificity, PPV, and NPV between malignant and benign lesions were 100%, 50.4%, 56.2%, and 100%, respectively. With a cut-off value of Ōēźcategory 4B, these values were 86.6%, 96.1%, 93.4%, and 91.1%, respectively (Table 1).
The AUC and cut-off value for each SWE parameter were assessed. The diagnostic performance of each SWE parameter to differentiate malignant from benign lesions are summarized in Table 2. Among qualitative and quantitative SWE parameters, a color pattern of 3 or 4 showed the highest sensitivity (92.7%) and NPV (94.2%). However, it had a low specificity (76.0%). Emean showed the highest specificity (86.8%), with a PPV of 77.9% and a low sensitivity of 73.2% (AUC, 0.859; cut-off>42.08 kPa). Emax showed the highest accuracy in differentiating malignant from benign lesions (AUC, 0.891; cut-off>50.85 kPa). Moreover, Emax had the highest AUC regardless of depth or size. In contrast, the color pattern was affected by the depth of the lesion (Table 3).
The associations between immunohistochemical factors and SWE parameters of the malignant tumors were analyzed and are shown in Table 4. Ki-67-positive lesions showed statistically significant differences in some parameters. Of the 60 Ki-67-positive lesions, 59 (98.3%) had a color pattern of 3 or 4, while 17 (77.3%) of the 22 Ki-67-negative lesions showed a color pattern of 3 or 4 (P=0.005). The SDmean of Ki-67-positive lesions was significantly higher than that of Ki-67-negative lesions (P=0.017). The Emean and Emax of Ki-67-positive lesions were higher than the corresponding values in Ki-67-negative lesions, although the differences were not statistically significant (P=0.090 and P=0.052, respectively). However, the Emax/Efat ratio of Ki-67-positive lesions was significantly higher than that of Ki-67-negative lesions (P=0.027). Poor tumor differentiation was correlated with higher SDmean (P=0.004) and Emax (P=0.021) values. PR negativity was also correlated with higher SDmean (P=0.003) and Emax (P=0.004) values, and HER2 positivity showed significant associations with higher SDmean and SDmax values (P=0.011 and P=0.023, respectively). CK5/6-positive tumors showed higher values of the Emax/Efat ratio than CK5/6-negative tumors (P=0.038). Luminal B, HER2-enriched, and triple-negative (non-basal) subtypes showed somewhat higher SDmean values than luminal A and triple-negative (basal) subtypes (P=0.028) according to ANOVA.
We found that Emax was the most predictive diagnostic parameter of SWE for differentiating malignant from benign lesions. In addition, SWE parameters were correlated with tumor differentiation and pathologic features related to tumor aggressiveness.
Many studies have revealed that quantitative SWE parameters can improve diagnostic performance, but each study used different cut-off values for the SWE parameters [21]. Park et al. [21] reported optimal cut-off values of 145.7 kPa and 89.1 kPa for Emax and Emean. Au et al. [22] reported optimal cut-off values of 46.7 kPa for Emax and 42.5 kPa for Emean. Chang et al. [15] used a cut-off value of 80.17 kPa for Emean in their study. Our cut-off values of quantitative SWE parameters for differentiating between benign and malignant lesions were 50.85 kPa and 42.08 kPa for Emax and Emean, respectively. The cut-off values derived in our data set are somewhat lower than those of other studies. The US system used in our study was different from the machine that previous studies [15,21,22] used, and in their studies, the SWE parameters of maximum, mean, and minimum stiffness values were derived by a single measurement at the stiffest portion of the mass. In our system, Emax values were measured at the stiffest portion of the mass using a fixed ROI, and the Emax and SDmax were the average and SD values of a fixed ROI. However, Emean and SDmean were measured by delicately drawing the ROI along the margin of the whole mass and then calculating the average elasticity value of the lesion and standard deviation. For this reason, evaluating Emax was simpler and took less time than assessing Emean. Moreover, our results showed that Emax was less affected by the depth or size of the lesion.
Our study investigated associations between immunohistochemical factors of breast cancer and various qualitative and quantitative parameters of SWE. Higher SDmean values were significantly correlated with poor tumor differentiation (P=0.004), Ki-67 positivity (P=0.017), PR negativity (P=0.003), and HER2 positivity (P=0.011). Furthermore, Ki-67-positive breast cancer was more likely to show a heterogeneous color distribution than Ki-67-negative breast cancer (P<0.05).
The Emax/Efat ratio was also significantly higher in Ki-67-positive and CK5/6-positive tumors. PR negativity was correlated with higher Emax (P=0.004). Moreover, the luminal B, HER2-enriched, and triple-negative (non-basal) subtypes showed somewhat high SDmean values than the luminal A and triple-negative (basal) subtypes (P=0.028), although the logistic regression analysis showed no significant difference. Thus, if the tumor has a heterogeneous shear wave and color distribution with a higher Emax value, we can predict that it is relatively likely to be poorly differentiated with a more aggressive immunohistochemical subtype, although a prospective validation study with a larger study population is needed for future research.
A high SDmean indicates internal elastic heterogeneity of the tumor. In other words, a more aggressive cancer not only becomes stiffer overall, but also has a heterogeneous distribution of stiffer regions within the tumor. In a fludeoxyglucose-positron emission tomography/computed tomography study using texture analysis, tumor heterogeneity was higher in invasive breast cancer with pathological factors predicting a poor prognosis [23]. This may be because more aggressive tumors exhibit various stages of histological heterogeneity, including proliferative tissue, hypoxic areas, and necrotic portions. Leek et al. [24] suggested that rapidly growing aggressive tumors have regions with hypoxic changes due to a lack of vascular supply, which leads to necrosis within the tumor [24].
This study has several imitations. First, our research was conducted using a relatively small sample size at a single center. Second, operator variability may have been a relevant factor, as in other US studies. For example, the fat lobule elasticity value was subjectively measured by three radiologists. In addition, there was no standardized location or method of measuring the elasticity value of fat the lobule. This could have resulted in some degree of inconsistency. To decrease operator variability, only expert breast radiologists with a 2-month learning period to achieve consensus for this SWE study evaluated the US examinations.
In conclusion, Emax is a relatively simple, reliable, and useful parameter for differentiating malignant from benign breast lesions. In addition, poor differentiation, PR-negative, and Ki-67-positive breast cancers were found to be associated with high stiffness and tumor heterogeneity. Thus, higher values of SDmean, Emax, and the Emax/Efat ratio could be used to predict these immunohistochemical factors. In such cases, more aggressive treatment plans can be considered.
AcknowledgementsStatistical consultation was supported by the Department of Biostatistics of the Catholic Research Coordinating Center.
NotesAuthor Contributions Conceptualization: Kang BJ. Data acquisition: Kang BJ, Lee J, Kim SH. Data analysis or interpretation: Kang BJ, Kim H. Drafting of the manuscript: Kang BJ, Kim H. Critical revision of the manuscript: Kang BJ, Kim H. Approval of the final version of the manuscript: all authors. References1. Taneja P, Maglic D, Kai F, Zhu S, Kendig RD, Fry EA, et al. Classical and novel prognostic markers for breast cancer and their clinical significance. Clin Med Insights Oncol 2010;4:15ŌĆō34.
2. Slamon DJ, Leyland-Jones B, Shak S, Fuchs H, Paton V, Bajamonde A, et al. Use of chemotherapy plus a monoclonal antibody against HER2 for metastatic breast cancer that overexpresses HER2. N Engl J Med 2001;344:783ŌĆō792.
3. Inwald EC, Klinkhammer-Schalke M, Hofstadter F, Zeman F, Koller M, Gerstenhauer M, et al. Ki-67 is a prognostic parameter in breast cancer patients: results of a large population-based cohort of a cancer registry. Breast Cancer Res Treat 2013;139:539ŌĆō552.
4. Jones RL, Salter J, A'Hern R, Nerurkar A, Parton M, Reis-Filho JS, et al. The prognostic significance of Ki67 before and after neoadjuvant chemotherapy in breast cancer. Breast Cancer Res Treat 2009;116:53ŌĆō68.
5. Yerushalmi R, Woods R, Ravdin PM, Hayes MM, Gelmon KA. Ki67 in breast cancer: prognostic and predictive potential. Lancet Oncol 2010;11:174ŌĆō183.
6. Nishimura R, Osako T, Okumura Y, Hayashi M, Arima N. Clinical significance of Ki-67 in neoadjuvant chemotherapy for primary breast cancer as a predictor for chemosensitivity and for prognosis. Breast Cancer 2010;17:269ŌĆō275.
8. Evans A, Whelehan P, Thomson K, McLean D, Brauer K, Purdie C, et al. Quantitative shear wave ultrasound elastography: initial experience in solid breast masses. Breast Cancer Res 2010;12:R104.
9. Cosgrove DO, Berg WA, Dore CJ, Skyba DM, Henry JP, Gay J, et al. Shear wave elastography for breast masses is highly reproducible. Eur Radiol 2012;22:1023ŌĆō1032.
10. Yeoh HJ, Kim TY, Ryu JA. The feasibility of shear wave elastography for diagnosing superficial benign soft tissue masses. Ultrasonography 2019;38:37ŌĆō43.
11. Youk JH, Gweon HM, Son EJ. Shear-wave elastography in breast ultrasonography: the state of the art. Ultrasonography 2017;36:300ŌĆō309.
12. Li G, Li DW, Fang YX, Song YJ, Deng ZJ, Gao J, et al. Performance of shear wave elastography for differentiation of benign and malignant solid breast masses. PLoS One 2013;8:e76322.
13. Ganau S, Andreu FJ, Escribano F, Martin A, Tortajada L, Villajos M, et al. Shear-wave elastography and immunohistochemical profiles in invasive breast cancer: evaluation of maximum and mean elasticity values. Eur J Radiol 2015;84:617ŌĆō622.
14. Youk JH, Gweon HM, Son EJ, Kim JA, Jeong J. Shear-wave elastography of invasive breast cancer: correlation between quantitative mean elasticity value and immunohistochemical profile. Breast Cancer Res Treat 2013;138:119ŌĆō126.
15. Chang JM, Park IA, Lee SH, Kim WH, Bae MS, Koo HR, et al. Stiffness of tumours measured by shear-wave elastography correlated with subtypes of breast cancer. Eur Radiol 2013;23:2450ŌĆō2458.
16. Denis M, Gregory A, Bayat M, Fazzio RT, Whaley DH, Ghosh K, et al. Correlating tumor stiffness with immunohistochemical subtypes of breast cancers: prognostic value of comb-push ultrasound shear elastography for differentiating luminal subtypes. PLoS One 2016;11:e0165003.
17. Tozaki M, Fukuma E. Pattern classification of ShearWave Elastography images for differential diagnosis between benign and malignant solid breast masses. Acta Radiol 2011;52:1069ŌĆō1075.
18. Jeong YS, Kang J, Lee J, Yoo TK, Kim SH, Lee A. Analysis of the molecular subtypes of preoperative core needle biopsy and surgical specimens in invasive breast cancer. J Pathol Transl Med 2020;54:87ŌĆō94.
19. Goldhirsch A, Winer EP, Coates AS, Gelber RD, Piccart-Gebhart M, Thurlimann B, et al. Personalizing the treatment of women with early breast cancer: highlights of the St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2013. Ann Oncol 2013;24:2206ŌĆō2223.
20. DeLong ER, DeLong DM, Clarke-Pearson DL. Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics 1988;44:837ŌĆō845.
21. Park J, Woo OH, Shin HS, Cho KR, Seo BK, Kang EY. Diagnostic performance and color overlay pattern in shear wave elastography (SWE) for palpable breast mass. Eur J Radiol 2015;84:1943ŌĆō1948.
22. Au FW, Ghai S, Moshonov H, Kahn H, Brennan C, Dua H, et al. Diagnostic performance of quantitative shear wave elastography in the evaluation of solid breast masses: determination of the most discriminatory parameter. AJR Am J Roentgenol 2014;203:W328ŌĆōW336.
A 63-year-old woman with a right breast mass (1-o'clock position, invasive ductal carcinoma, moderate differentiation, ER-, PR-, HER2-, Ki-67+: triple negative) underwent shear wave elastography.A. Transverse scan of ultrasonography shows a 2.3-cm oval hypoechoic mass with a microlobulated margin at 1-o'clock position of the right breast, suggesting category 4C (suspicious abnormality) (depth, 1.7 cm). B. The parameters of shear wave elastography are as follows: Color pattern=4, Emean=85.9, SDmean=32.5. C. The parameters of shear wave elastography are as follows: Emax, 137.1; SDmax, 27.3; Efat, 30.6; Emean/Efat, 2.8; Emax/Efat, 4.5. ER, estrogen receptor; PR, progesterone receptor; HER2, human epidermal growth factor receptor 2.
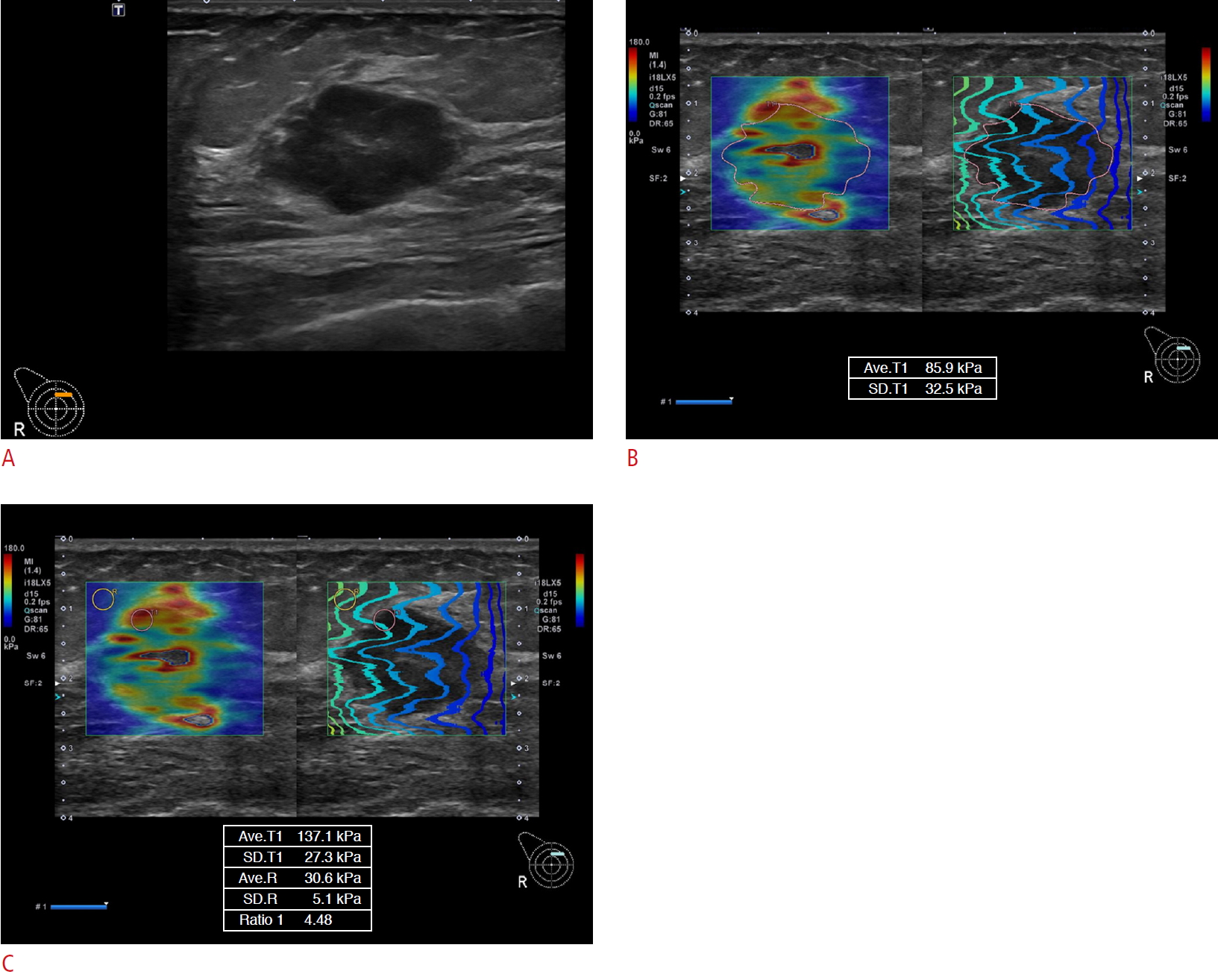 Fig.┬Ā1.A 51-year-old woman with a right breast mass (12-o'clock position, fibroadenoma) underwent shear wave elastography.A. Sagittal scan of ultrasonography shows a 1.2-cm oval hypoechoic mass with a partial angular margin in the 12-oŌĆÖclock position of the right breast, suggesting category 4A (suspicious abnormality) (depth=1.6 cm). B. The parameters of shear wave elastography are as follows: Color pattern, 1; Emean, 26.7; SDmean, 9.0. C. The parameters of shear wave elastography are as follows: Emax, 37.5; SDmax, 7.3; Efat, 16.9; Emean/Efat, 1.6; Emax/Efat, 2.2.
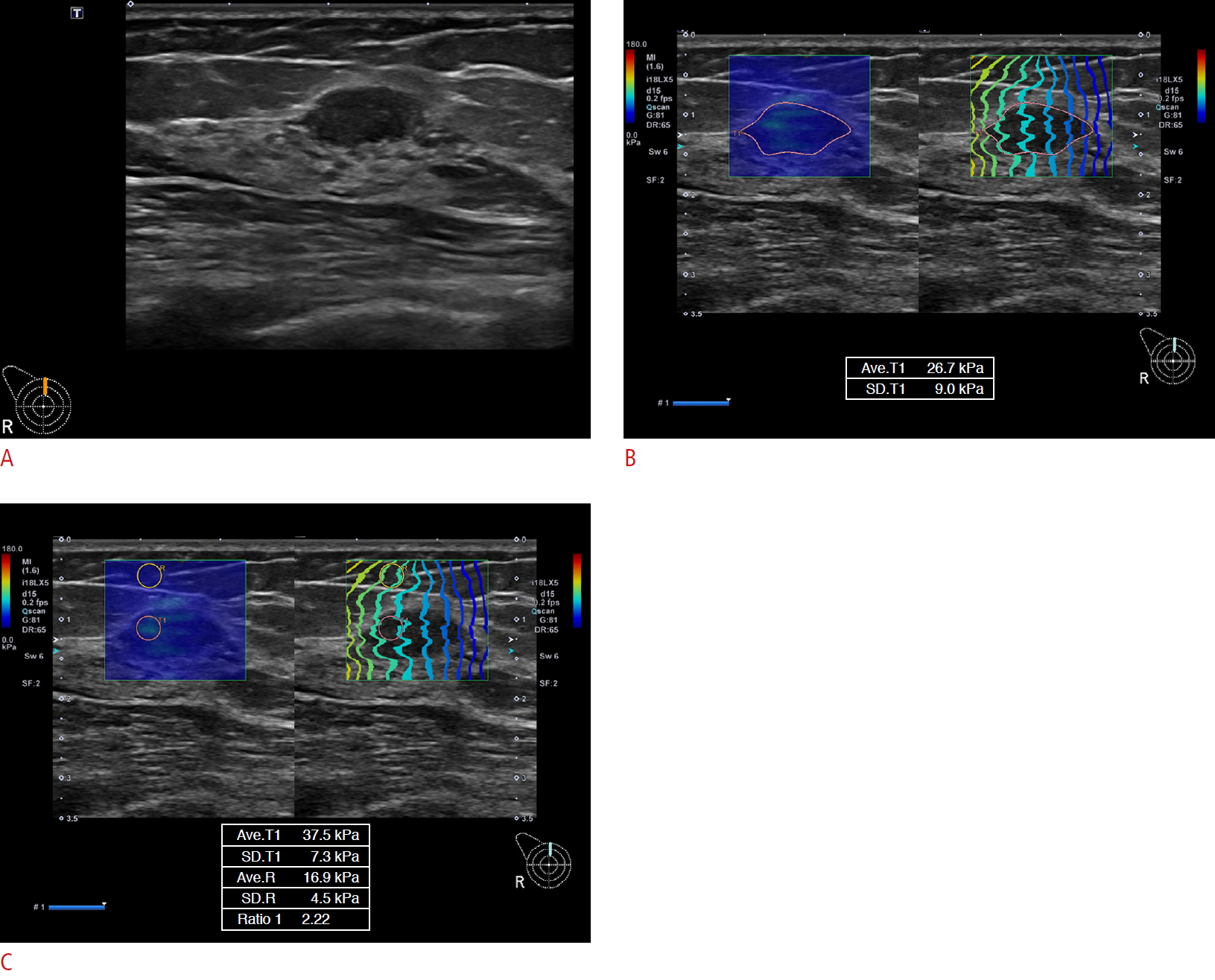 Fig.┬Ā2.Table┬Ā1.Baseline characteristics of the breast lesions Table┬Ā2.Diagnostic performance of qualitative and quantitative SWE parameters Table┬Ā3.AUC of SWE parameters according to category classification, depth, and size
Table┬Ā4.Associations between malignancy and immunopathologic factors
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||



 Print
Print facebook
facebook twitter
twitter Linkedin
Linkedin google+
google+ Download Citation
Download Citation PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC




